_67.png)
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
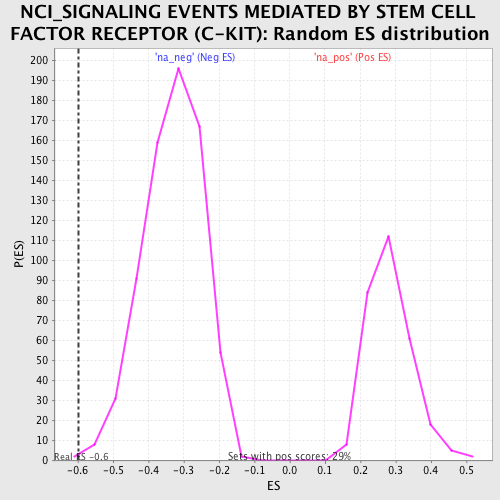
| GeneSet | NCI_SIGNALING EVENTS MEDIATED BY STEM CELL FACTOR RECEPTOR (C-KIT) |
| Enrichment Score (ES) | -0.59739816 |
| Normalized Enrichment Score (NES) | -1.8050936 |
| Nominal p-value | 0.0028169013 |
| FDR q-value | 0.12772933 |
| FWER p-Value | 0.872 |
_67.png)
| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GRB2 | 736 | 3.346 | -0.0126 | No | ||
| 2 | STAT5A | 1251 | 2.601 | -0.0192 | No | ||
| 3 | MATK | 1628 | 2.250 | -0.0212 | No | ||
| 4 | SNAI2 | 2163 | 1.920 | -0.0344 | No | ||
| 5 | PTPRO | 2500 | 1.755 | -0.0383 | No | ||
| 6 | MAP2K2 | 3298 | 1.440 | -0.0696 | No | ||
| 7 | MAPK8 | 3997 | 1.205 | -0.0974 | No | ||
| 8 | RPS6KB1 | 4288 | 1.133 | -0.1039 | No | ||
| 9 | EPO | 4950 | 0.969 | -0.1316 | No | ||
| 10 | RAF1 | 5344 | 0.875 | -0.1457 | No | ||
| 11 | LYN | 5405 | 0.860 | -0.1420 | No | ||
| 12 | PIK3R1 | 6233 | 0.693 | -0.1809 | No | ||
| 13 | GRAP2 | 6593 | 0.616 | -0.1953 | No | ||
| 14 | AKT1 | 6874 | 0.557 | -0.2058 | No | ||
| 15 | BAD | 7079 | 0.515 | -0.2126 | No | ||
| 16 | CREBBP | 7951 | 0.345 | -0.2568 | No | ||
| 17 | EPOR | 7953 | 0.345 | -0.2540 | No | ||
| 18 | DOK1 | 8858 | 0.179 | -0.3013 | No | ||
| 19 | HRAS | 9039 | 0.147 | -0.3098 | No | ||
| 20 | MAP2K1 | 9344 | 0.087 | -0.3254 | No | ||
| 21 | SPRED1 | 10327 | -0.132 | -0.3772 | No | ||
| 22 | SOS1 | 10560 | -0.179 | -0.3883 | No | ||
| 23 | KIT | 11438 | -0.360 | -0.4326 | No | ||
| 24 | GRB10 | 11447 | -0.363 | -0.4301 | No | ||
| 25 | MITF | 13644 | -0.915 | -0.5410 | No | ||
| 26 | MAPK3 | 13948 | -1.009 | -0.5491 | No | ||
| 27 | FOXO3 | 14022 | -1.039 | -0.5446 | No | ||
| 28 | PIK3CA | 14498 | -1.205 | -0.5605 | No | ||
| 29 | PTPN11 | 15185 | -1.479 | -0.5854 | Yes | ||
| 30 | SH2B3 | 15399 | -1.593 | -0.5840 | Yes | ||
| 31 | JAK2 | 15588 | -1.691 | -0.5804 | Yes | ||
| 32 | PTEN | 15842 | -1.843 | -0.5791 | Yes | ||
| 33 | PTPN6 | 15968 | -1.915 | -0.5704 | Yes | ||
| 34 | GSK3B | 16087 | -1.994 | -0.5606 | Yes | ||
| 35 | KITLG | 16112 | -2.015 | -0.5456 | Yes | ||
| 36 | MAP4K1 | 16285 | -2.168 | -0.5373 | Yes | ||
| 37 | SHC1 | 16333 | -2.217 | -0.5219 | Yes | ||
| 38 | SOCS1 | 17323 | -3.435 | -0.5473 | Yes | ||
| 39 | CBL | 17607 | -4.019 | -0.5300 | Yes | ||
| 40 | CRKL | 17893 | -4.821 | -0.5064 | Yes | ||
| 41 | STAT3 | 17907 | -4.850 | -0.4678 | Yes | ||
| 42 | VAV1 | 17943 | -4.944 | -0.4297 | Yes | ||
| 43 | PDPK1 | 18064 | -5.362 | -0.3927 | Yes | ||
| 44 | SPRED2 | 18094 | -5.476 | -0.3500 | Yes | ||
| 45 | STAT1 | 18476 | -8.708 | -0.3000 | Yes | ||
| 46 | GAB1 | 18529 | -10.215 | -0.2201 | Yes | ||
| 47 | BCL2 | 18536 | -10.511 | -0.1354 | Yes | ||
| 48 | TEC | 18590 | -17.252 | 0.0014 | Yes |